It is widely known that defining and diagnosing mental disorders is a difficult process due to overlapping nature of symptoms, and lack of a biological test that can serve as a definite and quantified gold standard. Mental Disorders such as Attention Deficit Hyperactivity Disorder (ADHD) and Autism Spectrum Disorder (ASD) are notoriously difficult to diagnose, especially in children. The current psychiatric diagnostic process is based purely on the behavioral observation of symptomology (DSM-5/ICD-10) and may be prone to misdiagnosis. The current psychiatric diagnosis is based purely on behavioural observation (DSM-5/ICD-10) but lacks biological and/or genetic validity and is prone to errors. For most mental health disorders definitive and quantitative diagnostic tests which can detect the presence or the absence of a specific psychiatric disorder(s) are non-existent.
Functional Magnetic Resonance Imaging (fMRI) is a non-invasive technique for studying the brain functional activities and is based on Blood Oxygen Level Dependent (BOLD) contrast. During the fMRI scan, a series of images are taken using a scanner while the subject to a specific task such as resting, or doing various pre-determined tasks. The result of brain scanning is a series of low resolution images over time which shows the activity of the brain and can allow us to develop a highly informative brain connectome. There is evidence that ADHD, Bipolar Disorder (BD) and Schizophrenia have characteristics that differ in the regional and global connectivity of the brain when studied under resting state Functional Magnetic Resonance Imaging or fMRI.
As computational scientists, along with our colleagues from psychiatry and neuroscience, the overarching question that we have been posing is as follows: Can we diagnose a person with ASD and ADHD (or other mental disorders) using fMRI scans using techniques from machine learning and novel algorithmic designs? If possible, such a test will provide a definitive quantitative ’gold standard’ test for diagnosing ADHD and other disorders.
In this endeavor machine-learning, especially deep-learning algorithms, have the potential to show exceptional promise. To this end, we have been successful in developing a machine learning algorithm that allow us to classify fMRI ADHD scans from normal healthy brain scans without using any demographic information. Our recently proposed technique is based on computing similarity between two multivariate time series along with k-Nearest-Neighbor classifier. We designed a model selection scheme called J-Eros which is able to pick the optimum value of k for k-Nearest-Neighbor from the training data. Our results show a 20% increase in accuracy, with superior sensitivity and specificity, as compared to the state of the art algorithms in classifying ADHD using open-data ADHD-200 that is available to the community.
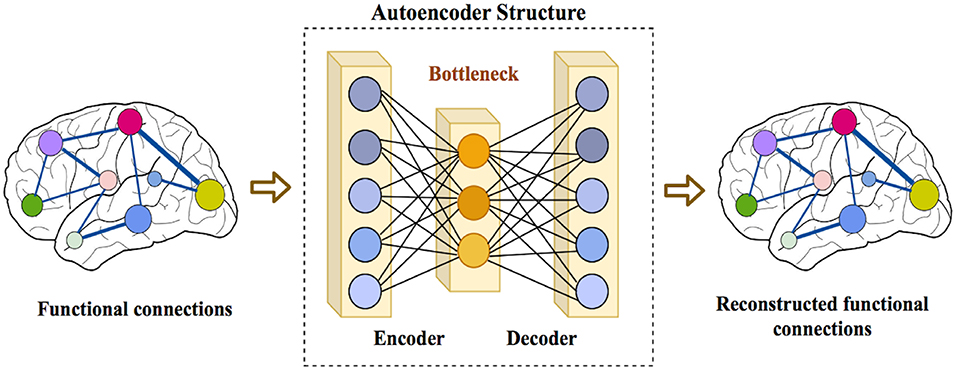
We also proposed a framework called ASD-DiagNet for classifying subjects with ASD from healthy subjects by using only fMRI data. We designed and implemented a joint learning procedure using an autoencoder and a single layer perceptron (SLP) which results in improved quality of extracted features and optimized parameters for the model. Further, we designed and implemented a data augmentation strategy, based on linear interpolation on available feature vectors, that allows us to produce synthetic datasets needed for training of machine learning models. The proposed approach is evaluated on a public dataset provided by Autism Brain Imaging Data Exchange including 1,035 subjects coming from 17 different brain imaging centers. Our machine learning model outperforms other state of the art methods from 10 imaging centers with increase in classification accuracy up to 28% with maximum accuracy of 82%. The machine learning technique presented in this paper, in addition to yielding better quality, gives enormous advantages in terms of execution time (40 min vs. 7 h on other methods).
Compute time for analyzing big fMRI data is a bottleneck in introducing such techniques in a clinical setting. High performance computing (HPC) techniques will help ease the computational load in big fMRI data analysis for clinical and diagnostic purposes. Among parallel computing techniques, Graphic Processing Unit (GPU) architecture provides a high degree of parallelism while being less expensive. Therefore, the focus of our research has been towards designing GPU based parallel computing algorithms for analyzing big fMRI data and considering pairwise relations between all voxels (the smallest cubic unit in fMRI data) without spatial or temporal constraint. In this way, voxels from different regions of brain which have similar spatiotemporal behaviour could be placed in the same cluster. The resulting clusters are regions which are functionally homogeneous and would give a more system-wide view of brain connectivity. This will allow neuroscientists to study the regional and global connectivity of the brain in much fine-grained detail than is currently possible. To this end, we have recently presented a GPU based strategy that allows us to calculate the functional connectivity using Pearson correlations for big fMRI data.
Such quantified psychiatric evaluations based on data science, big data, and algorithms can increase the accuracy of diagnosis, prognosis, and treatments of difficult to assess mental disorders. It will be an immense service for mankind if we are successful in developing a quantitative framework for big fMRI data to identify brain disorders from healthy brains. These algorithms will play a vital role in peering into mental disorders and will be akin to the role that microscope played for diagnostic medicine in the early 19th century.
Recent Publications
- Taban Eslami, Joseph S Raiker, Fahad Saeed, “Explainable and Scalable Machine-Learning Algorithms for Detection of Autism Spectrum Disorder using fMRI Data”, Book Chapter In Neural Engineering Techniques for Autism Spectrum Disorder, pp. 39-54. Academic Press, 2021 Elsiever | arXiv
- Taban Eslami, Fahad Almuqhim, Joseph S. Raiker, Fahad Saeed*, “Machine Learning Methods for Diagnosing Autism Spectrum Disorder and Attention- Deficit/Hyperactivity Disorder Using Functional and Structural MRI: A Survey“, Frontiers of Neuroinformatics, Vol. 14, pp 62, 2021 Frontiers
- Fahad Almuqhim, and Fahad Saeed*, “ASD-SAENet: Sparse Autoencoder for detecting Autism Spectrum Disorder (ASD) using fMRI data“, Frontiers in Computational Neuroscience, Vol. 15, p. 27, March 2021 Frontiers
- Taban Eslami, Vahid Mirjalili, Alvis Fong, Angela R. Laird, and Fahad Saeed*, “ASD-DiagNet: A Hybrid Learning Approach for Detection of Autism Spectrum Disorder Using fMRI Data” Frontiers of Neuroinformatics, vol. 13, pages 70, Nov 2019 arXiv | Frontiers (Best Paper Editor’s Pick 2021)
- Taban Eslami, and Fahad Saeed*, “Auto-ASD-Network: A technique based on Deep Learning and Support Vector Machines for diagnosing Autism Spectrum Disorder using fMRI data”, Workshop on Machine Learning Models for Multi-omics Data Integration, in Proceedings of 10th ACM Conference on Bioinformatics, Computational Biology (ACM BCB), Niagara Falls, New York, September 7-10, 2019. ACM | Tech Report
- Taban Eslami, and Fahad Saeed*, “Fast-GPU-PCC: A GPU-Based Technique to Compute Pairwise Pearson’s Correlation Coefficients for Time Series Data – An fMRI Study“, MDPI High-Throughput, April 2018 MDPI | PubMed
- Taban Eslami, and Fahad Saeed*, “GPU-DFC: A GPU-based parallel algorithm for computing dynamic-functional connectivity of big fMRI data“, Proceedings of IEEE International Conference On Big Data Service And Applications (IEEE Big Data Service 2019), San Francisco East Bay, California, USA, April 4 – 9, 2019 IEEE Xplore | Tech Report
- Taban Eslami, and Fahad Saeed*, “Similarity based classification of ADHD using Singular Value Decomposition“, Proceedings of ACM Conference on Computing Frontiers (ACM-CF), Ischia, Italy, May 2018 ACM | Tech Report | Presentation (YouTube)
