The Saeed Lab develops machine-learning models, combined with high-performance computing, and data science approaches, to study the functional genomics and organization of the human brain.
Our work focuses on understanding the stochastic difference between identified peptides from high-throughput mass spectrometry data for applications related to human health, disease, and environment. In addition, our work focuses on understanding brain function in the context of prediction, diagnosis and characterization of biomarkers specific to disorders such as epilepsy, ADHD, Autism, and Alzheimer’s. Improved computational methods may impact detection of novel biomarkers and non-abundant proteins, and better diagnosis/prediction outcomes for the patients.
To achieve these goals, we embrace open science principles and adopt and develop best practices to promote reproducible computational results. If you would like to know more about specific projects, you are welcome to visit us on GitHub and Software pages.
Current Projects
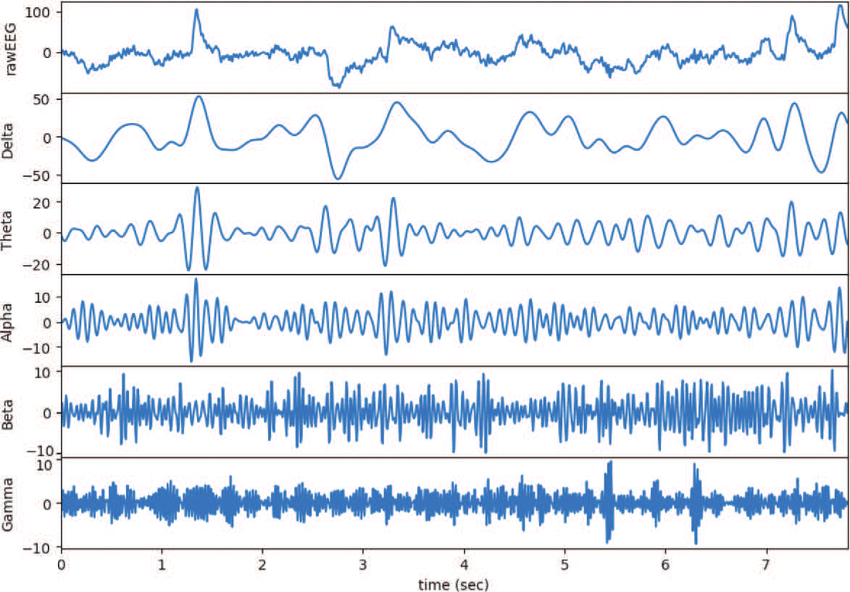
Predicting Epileptic Seizures
We investigate deep-learning methods to predict epileptic seizures before they happen.
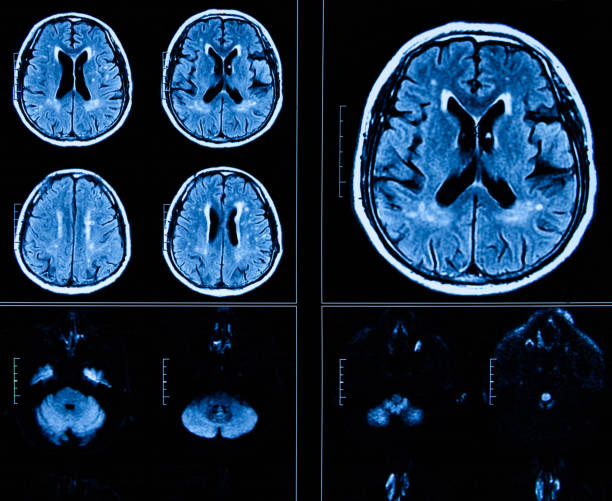
Predicting onset of Alzheimer’s
We investigate neuroimaging and machine-learning models to predict Alzheimer’s before any onset of symptoms.
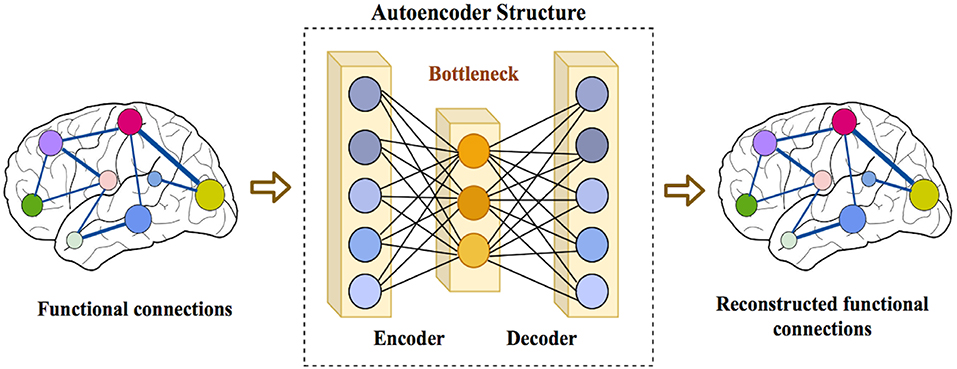
Characterization of Autism Spectrum
We investigate machine-learning models to classify and characterize the hetregenoity of autistic spectrum
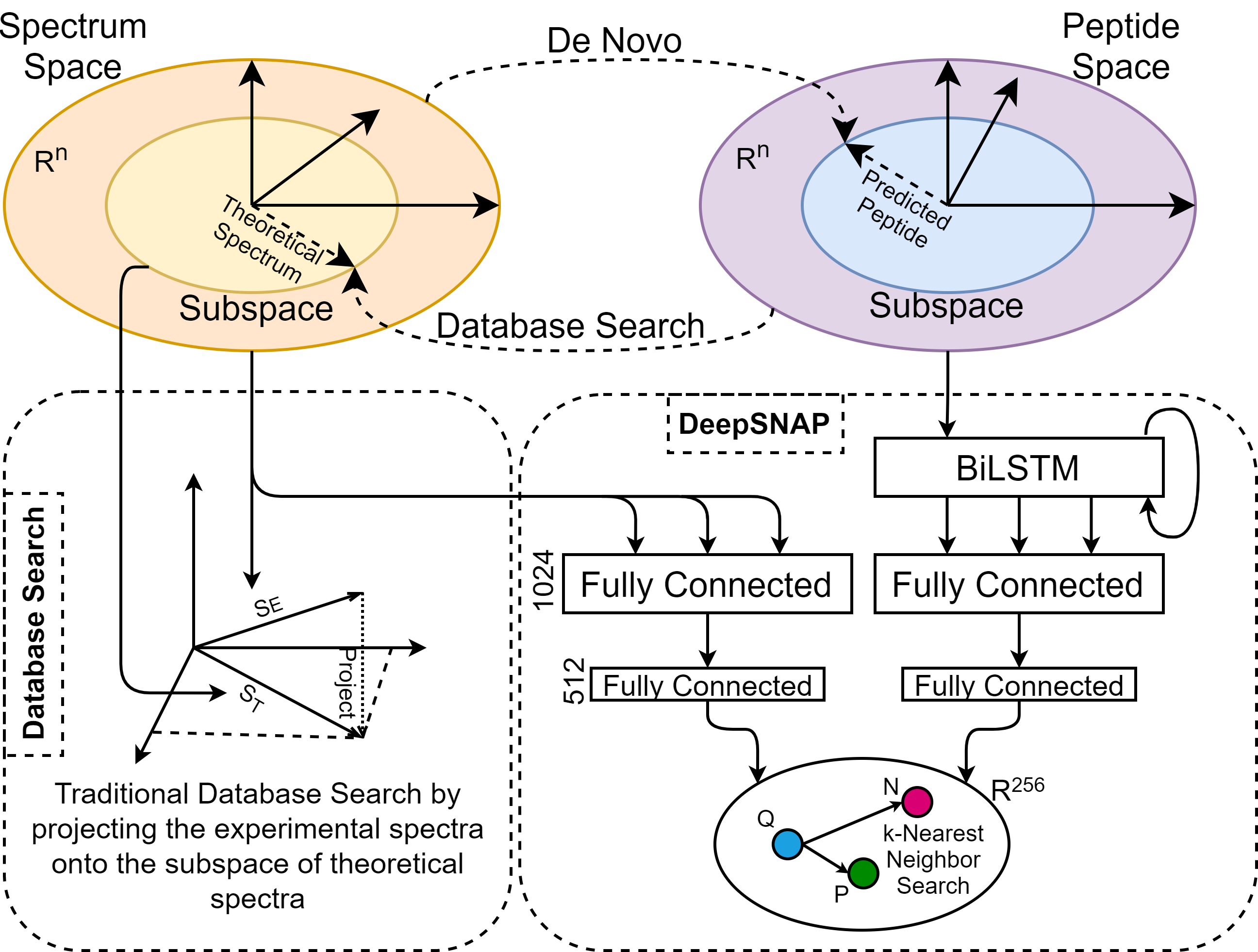
ML Ecosystem for Mass Spectrometry Data
An develop an interconnected set of open-source machine-learning tools for mass spectrometry based omics

